Quick Introduction to the Workshop and Core
The mission of the Bioinformatics Core facility is to facilitate outstanding omics-scale research through these activities:

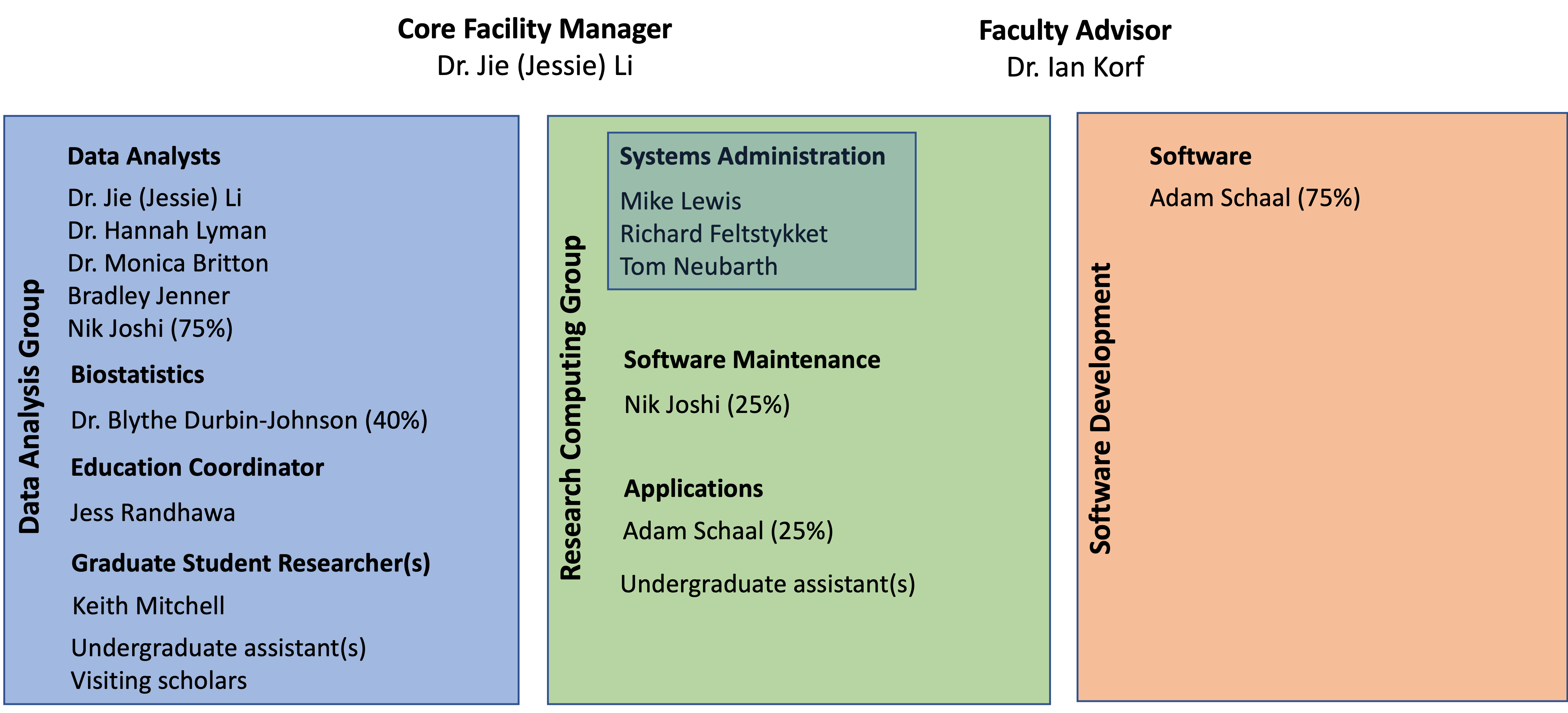
Staff and Students
Our team offers custom bioinformatics services to academic and private organizations. We have a strong academic background with a focus on cutting edge, open source software. We replicate standard analysis pipelines (best practices) when appropriate, and/or develop novel applications and pipelines when needed, however we always emphasize biological interpretation of the data.
Contacts
-
Request for data analysis services, consultations.
-
Bioinformatics training course information
-
Computing Issues, include but not limited to user account questions, equipment failure/malfunction, software install, software failures (not related to use).
Workshop Goals
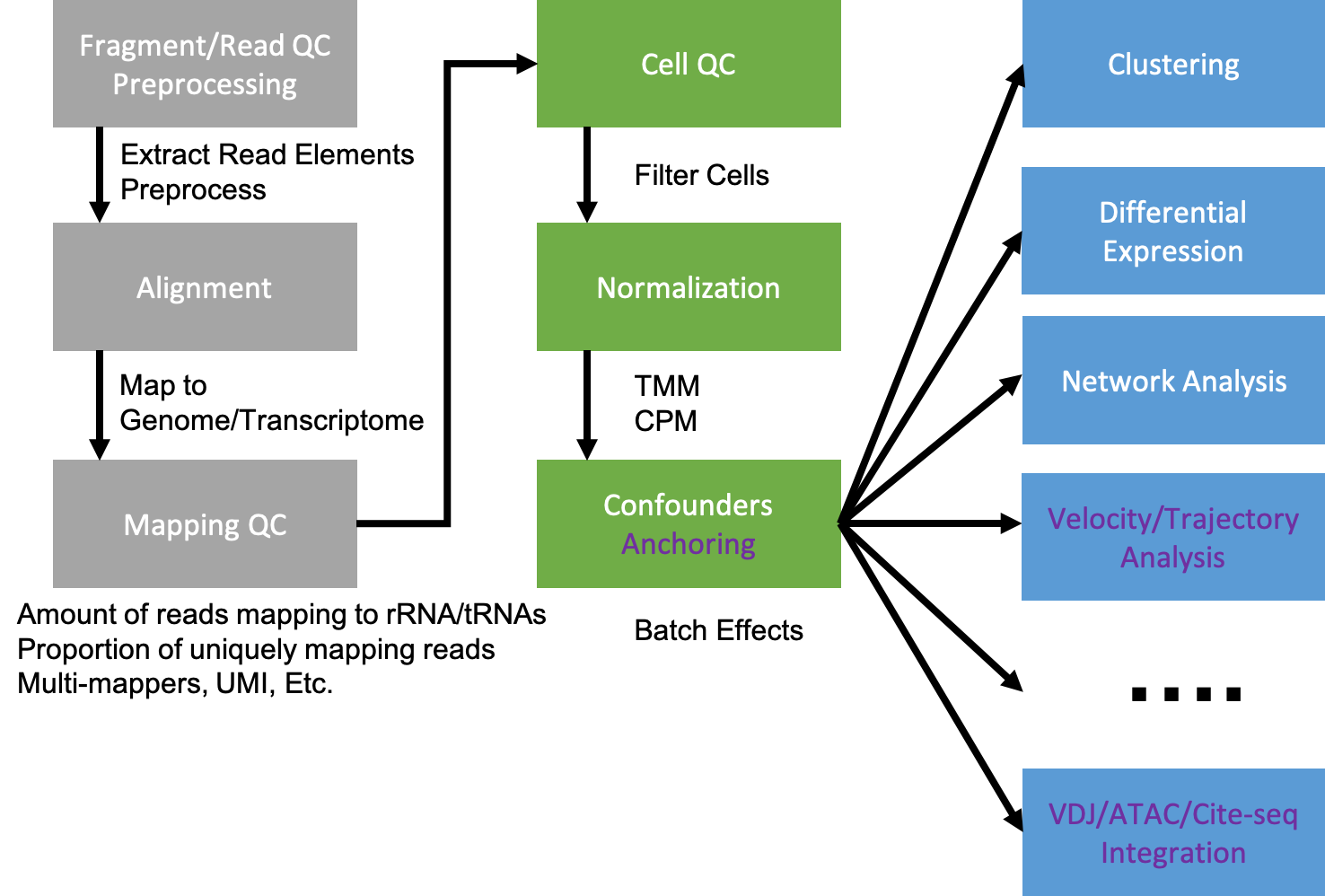
This workshop focuses on velocity and trajectory analysis. For differential expression analysis, consider Single Cell RNA-Seq Analysis.
Structure of the Virtual Workshops
Given the current Covid19 pandemic, and stay in place orders around the world, we’ve decided to take our popular bioinformatic workshops virtual. Our intent is to offer as close to an experience we can to our in-person workshops. We will hold the same goals and strive for a similar lecture/hand-on ratio. We will be using multiple technologies in order to help facilitate a maximum amount of interaction.
Zoom
Course lecture, discussions, and one-on-one help/troubleshooting will be conducted using a zoom meeting.
- We will be using breakout rooms for group work, one-on-one troubleshooting by our staff, screen sharing and even remote control.
- The Chat features of Zoom will not be our primary mode of text communication as its history is not reliable.
- Video recordings of lectures will be made available to participants.
Because video is involved, we ask everyone to be respectful and we reserve the right to remove someone if they are being disrespectful or disruptive.
Learn more about how we use Zoom in our workshops.
Slack
Text based communication will be conducted via a Slack channel. Staff will be monitoring the Slack channel to answer questions (and schedule a Zoom break out room if needed). If you know the answer to someone else’s question, feel free to answer it.
Learn more about how we use Slack in our workshops.
Workshop Materials
Workshop materials are all posted on github, and publicly available
http://bioinformatics.ucdavis.edu/training
-
Workshop registration site:
-
Public Github repository listing:
-
This Advanced scRNAseq Workshop
Computing Cluster
A portion of this course will be conducted on our servers and compute cluster (tadpole.genomecenter.ucdavis.edu).
Instructions on how to get an account will be sent by email.
